Q-Chem
Overview
Q-Chem is a comprehensive ab initio quantum chemistry program. Q-Chem’s capabilities facilitate applications in pharmaceuticals, materials science, biochemistry and other fields.
Q-Chem 6.3 Shared Memory and GPU Versions are available.
Restrictions on use
Q-Chem 6.3 is limited to one research group.
License Terms & Conditions of Q-Chem can be seen here: https://q-chem.com/purchase/license/
To get access to Q-Chem on the CSF3, you must request it via our help form and confirm that you have read the above License Terms & Conditions and your usage will comply with the Terms & Conditions of the License. On getting your request along with your confirmation and after approval of the license holder, you will be added to a secondary unix group which will enable your access to Q-Chem software in CSF3.
Set up procedure
To use Q-Chem load one of the following modulefiles:
module load apps/binapps/qchem/6.3 # For Serial and SMP version module load apps/binapps/qchem/6.3_gpu # For GPU Version
You do not need to load CUDA modules separately.
Running the application in CSF3
Please note that IQmol is not available in CSF3. If you need to use it for creating your input files, you can download and install IQmol in your Laptop/Desktop, it is available for free. For Windows there is a portable (ZIP) version available which can run without installing. After preparing input files in your Laptop/Desktop using IQmol, you can then upload the input file to CSF3 and submit it to the batch system using a suitable jobscript. Example jobscripts are given below.
Instructions on running batch jobs in CSF3 using IQmol software installed in your laptop/desktop can be seen below in the section titled Running the application in CSF3 using IQmol software in your laptop/desktop
A scratch directory named “qchemscratch” will be created within the job directory by Q-Chem when the job runs. It is used for storing temporary files and for restarting jobs and is emptied by Q-Chem on completion of the job.
$HOME/.qchemrc file.
Alternately, this can be also be done by setting those values in the jobscript. For example adding the line: export MEM_TOTAL="6000" after the module load lines will override the global MEM_TOTAL value of 4000MB to 6000MB. Variable MEM_TOTAL sets the amount of memory per-core that can be used by qchem. Please refer the maximum memory available in different node types before overriding the global defaults. Global defaults for SERIAL and SMP are set as 4000MB per core and for GPU it is set as 10000MB per core.
Serial job
An example batch submission script that will read a file named input.in from the current directory and write the output in input.out:
#!/bin/bash --login #SBATCH -p serial # (or --partition=) Use the nodes dedicated to 1-core jobs #SBATCH -t 1-0 # 1 day wallclock (max permitted is 7 days) module purge module load apps/binapps/qchem/6.3 qchem input.in input.out
Submit with the command: sbatch scriptname where scriptname is the name of your jobscript.
SMP Parallel job – 2 to 168 cores
An example batch submission script that will read a file named input.in from the current directory and write the output in input.out:
#!/bin/bash --login #SBATCH -p multicore # (or --partition=) Use the AMD 168-core jobs #SBATCH -n 4 # (or --ntasks=) Number of cores #SBATCH -t 1-0 # 1 day wallclock (max permitted is 7 days) module purge module load apps/binapps/qchem/6.3 qchem -nt $SLURM_NTASKS input.in input.out
Submit with the command sbatch scriptname where scriptname is the name of your jobscript.
GPU job
An example batch submission script that will read a file named input.in from the current directory and write the output in input.out:
#!/bin/bash --login #SBATCH -p gpuL # L40S GPUs. Other GPUs available: gpuA (A100-80GB) #SBATCH -G 1 # (or --gpus=) Number of GPUs #SBATCH -n 4 # (or --ntasks=) Number of cores. Up to 12 CPU cores per GPU permitted for L40S. #SBATCH -t 1-0 # 1 day wallclock (max permitted is 7 days) module purge module load apps/binapps/qchem/6.3_gpu qchem -gpu -nt $SLURM_NTASKS input.in input.out
Submit with the command sbatch scriptname where scriptname is the name of your jobscript.
NOTE: You don’t need to load any CUDA module separately.
Running the application in CSF3 using IQmol software in your laptop/desktop
You can install IQmol software in your laptop/desktop and use it to set up and submit your Q-Chem jobs directly to CSF3 batch system. You will need to configure the IQmol software after installing it to be able to submit jobs to CSF3. Please refer the below screenshots to set it up correctly:
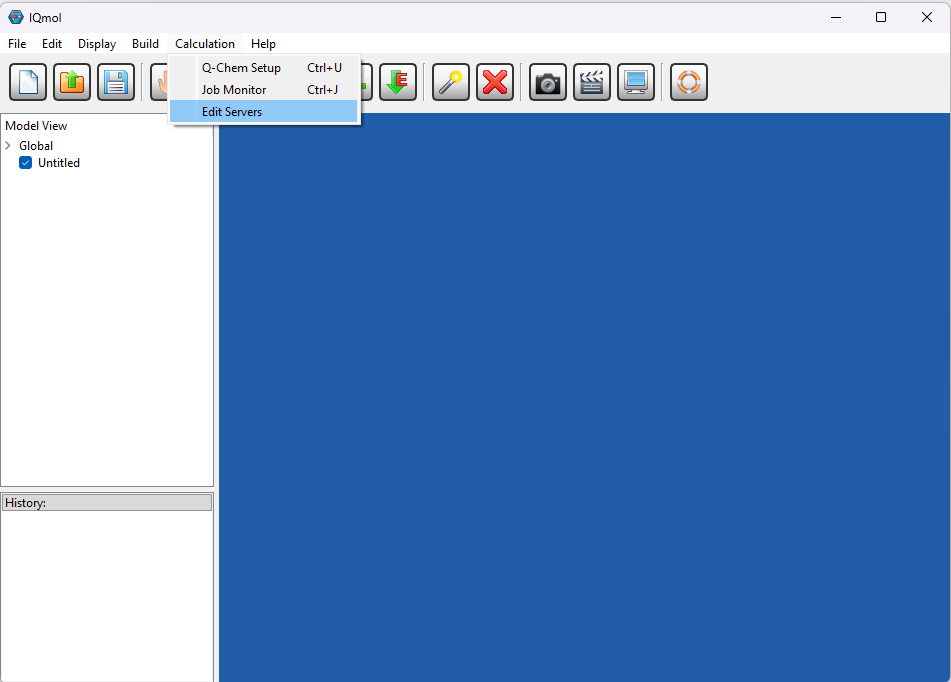
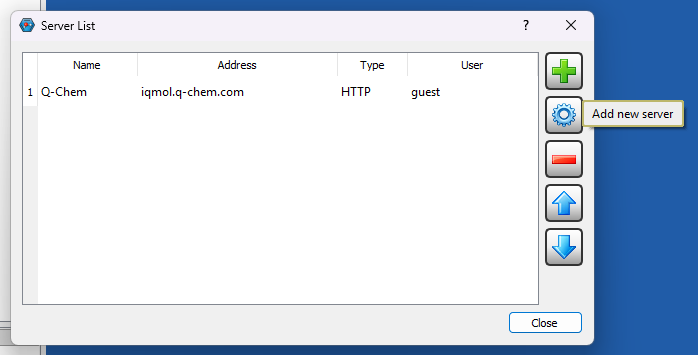
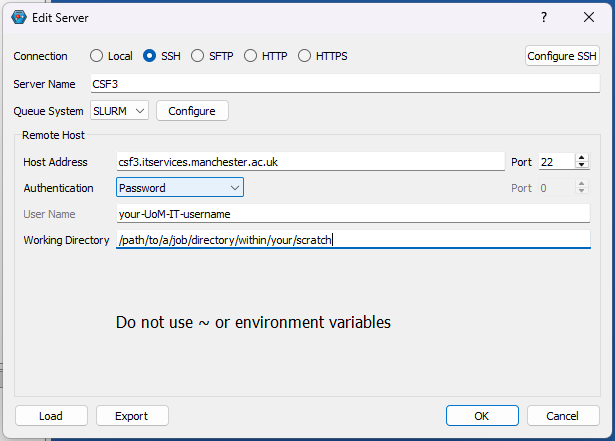
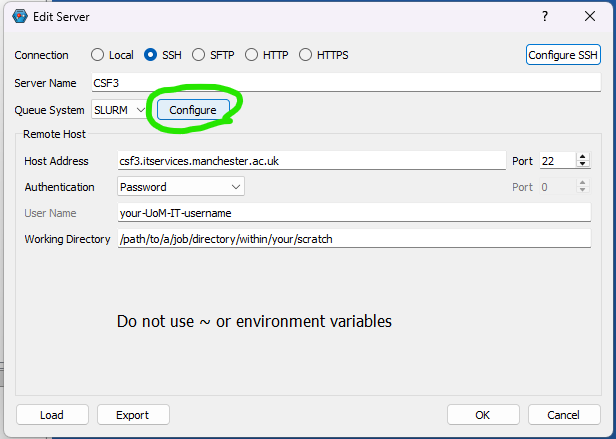
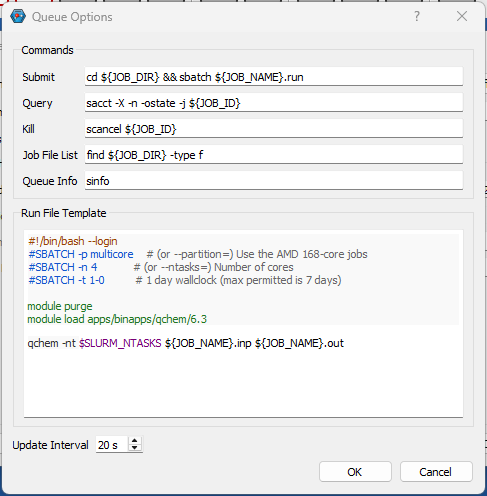
You can set up multiple instances of CSF3 server for different type of jobs (Serial/SMP/GPU), just add appropriate jobscript lines for them as shown above referring the sample jobscripts provided for them.
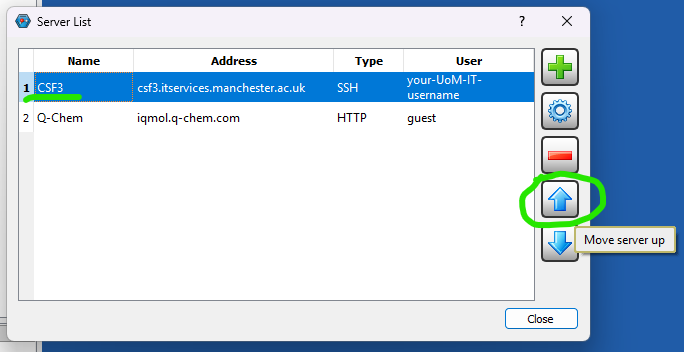
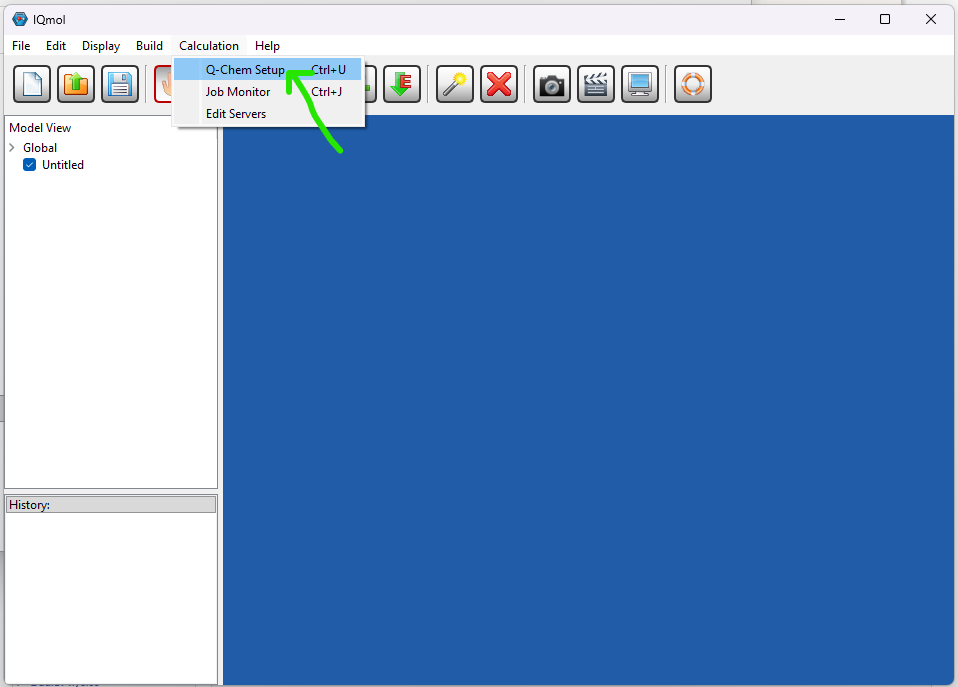
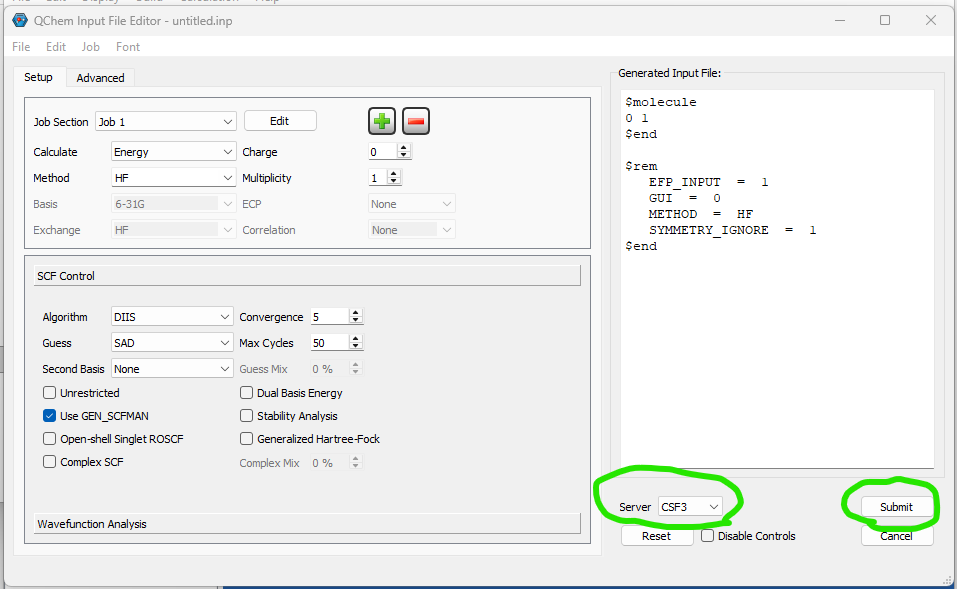
Further info
Updates
None at this time.
